| Fe
| Ti
| Ta1
| Ta2
| Ta3
| Ta4
| Ta5
| LR
| F/T
| BR
|
PI
| 1920
| 1910
| 2340
| 1200
| 910
| 760
| 380
| 1.79-2.27*
| 1.02-1.03
| 1.30-1.46
|
PII
| 2030
| 2030
| 880
| 540
| 430
| 330
| 250
| 0.44-0.45
| 0.98-1.01
| -
|
PIII
| 2300
| 2350
| 1010
| 700
| 590
| 380
| 280
| 0.43-0.51
| 0.97-0.99
| -
|
*these figures do not seem to be compatible with their data in Table 4 Very similar to C. pallidivittatus, but males can be differentiated by characters of the hypopygium, viz. Dististyle and inferior volsella longer and more tapered, superior volsella longer, anal point broader, and indentation in tergite IX is more a U-shape.
Additional data including further specimens:
Wing length abt 4.55-5.40 mm, width 1.14-1.32; VR 0.95; 4 Scf on brachiolum; 25-41 setae on squamal fringe.
Head: AR 3.34 (2.72-3.65); CT 29.7 (21-35) µm long and 1.90-2.0 longer than wide; 61-65 setae on clypeus, which is about 0.85-0.94 of the diameter of the antennal pedicel; palp proportions (µm) 82 : 75 : 278 : 273 : 328 ; P5/P4 1.15-1.33, P5/P3 1.23-1.56.
Thoracic setae: Acrostichal - abt 12-16; Dorsocentrals 32-36; Prealars - 1; Supraalars 6-9; Scutellar 8-20 in anterior rows, 11-22 in posterior row (total 22-42).
Leg lengths (micron) and proportions:
| Fe
| Ti
| Ta1
| Ta2
| Ta3
| Ta4
| Ta5
| LR
| F/T
| BR
|
PI
| 2104
| 2074
| 2418
| 1259
| 994
| 828
| 420
| 1.14-1.32
| 1.01-1.02
| 1.10-1.48
|
PII
| 2410
| 2142
| 995
| 752
| 460
| 357
| 272
| 0.44-0.47
| 0.98-1.01
| -
|
PIII
| 2400
| 2483
| 1242
| 757
| 626
| 424
| 303
| 0.43-0.56
| 0.94-0.99
| -
|
One specimen had 3 setae in a small clear area right at the base of the anal point. Inferior Volsella reaching to about 2/3 the length of the gonostylus, with simple setae along the distal 2/3. Gonostylus narrows gently along its length usually to more of a point than figured by Shobanov et al. (1999).Female: The female has not been described, but it is likely that the genital regions will be quite similar to those of C. tentans as described by Shilova (1978).
Larva described by Johannsen (1938). Chromosomes have been shown in a number of papers, e.g. Thompson (1971), Firling and Kobilka (1979), Martin (1979); full karyotype, using Beermann maps, published by Kiknadze et al. (1996a). Keyl (1962) gives the sequence of arm F on his scheme, including IR-2 = p'F3 = n'F1 (Kiknadze et al. 1996a). Other sequences from Kiknadze et al. (2004).
Go to Immatures and Cytology.
Found: Numerous localities across the northern U.S. and Canada.-
Alberta (WR) - Elkwater; Edmonton; Lacombe.
British Columbia (WC) - Chilcotin area; Williams Lake, Kamloops; Sawmill Lake, Westwick Lake (52.00°N; -122.17°W); Lake Sorenson (A.B. Acton).
Manitoba (WR) - Erickson; St. Alphonse; Churchill.
Ontario (ER) - Hogs Back (45.35oN, -75.68oW), Ottawa; Kars Creek, Kars (45.13°N, -75.63°W); Mud Lake, South March
(45.40°N, -75.87°W), all Carleton Co.; Cambridge (43.40°N, -80.30°W) (Telfer et al. 2015); Kelly Lake, Sudbury
(46.45°N, -81.07°W); Toronto.
Quebec (ER) - Montreal.
Saskatchewan (WR) - Big Quill Lake, 1 ml. s. Dafoe (51.73°N, -104 53°W); 6ml. n. & e. Colgate; Lake Waskesiu,
Prince Albert Park; 6ml. s. & w. Stoughton; 5ml. w. Theodore (51.75°N, -102.95°W).
Colorado - Walden Pond, Boulder.
Iowa (ER) - Lake Okoboji & Jemmerson Slough, Dickinson Co.; Little Wall Lake, Hamilton Co.;
Cerro, Gordo Co.
Massachusetts (ER) - Longmeadow
Michigan (ER)
Minnesota (WR) - Badger Lake, Erskine, Polk Co. (47.67°N, -96.00°W); Lake Christina, Douglas Co.
New Mexico - Taos, Taos County (36.394°N, 105.577°W).
New York (ER) - Ithaca.
North Dakota (WR) - Warsing Dam, Eddy Co.(47.83°N, -99.12°W); Braddock Dam, Emmons Co.; Fullers Slough;
Hankinson.
South Dakota (WR) - Lake Francis Case, Gregory Co. (43.27°N, -99.00°W); Gavins Point National Fish Hatchery,
(42.87°N, -97.47°W); 4 ml N Yankton, both Yankton Co.; Su-Falls; Missouri R., Running Water.
Utah (WR) - Logan, Cache Co.
Wisconsin (ER) - Stevens Pond & U.W. Arboretum, Madison (43.03°N, -89.42°W);, Dane Co.; Theresa Marsh.
Wyoming (WR) - 6ml s Lander.
In prairie sloughs, shallow eutrophic lakes and ponds, sewage oxidation lagoons.
Acton and Scudder (1971) consider North American populations to comprise three races - Alaskan, west Canadian, east Canadian. Alaskan is considered here to be still probably C. tentans (see Sp3y).
The other two races are distinct in the West and East respectively, but tend to merge in the central area where the biogeographic barriers currently exist, although Gunderina et al. (1996) showed that samples from Minnesota and Saskatchewan were distinct from the eastern populations. However this could just reflect an east-west cline. Therefore inversion frequency data from all available populations were incorporated in a UPGMA analysis (see below), which still supported two groupings, with a break somewhere around the western borders of Wisconsin in the U.S.A., and Ontario in Canada.
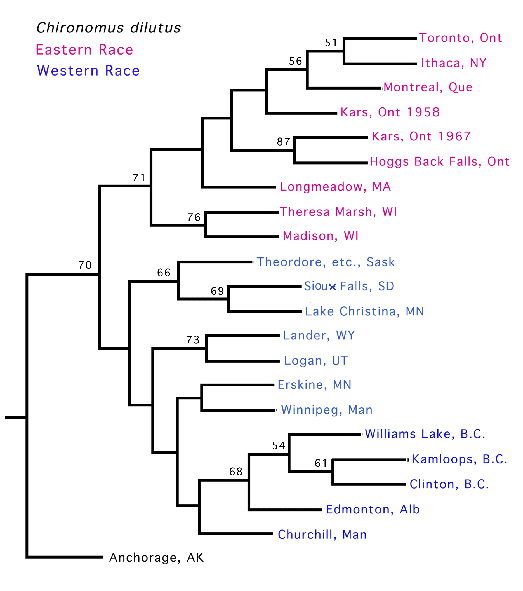
UPGMA tree showing the distribution of the two races of C. dilutusEastern Race: Larval length - female 24.5 mm (21.5-28.9)(4); male 20.0 mm (16.9-24.2)(3).
Cytologically, this race is characterized by more frequent n'D2, h'E2, and n'F4, but lower frequencies of n'F3 and n'G3; with no n'G4 recorded.
Western Race: Larval length - female 22.5 mm (19.8-27.0)(13); male 19.1 mm (15.3-22.5)(14).
Cytologically, this race is characterized by more frequent n'F3, n'G3 and rarely n'G4 (known only from British Columbia), while frequencis of n'D2 and h'E2 are lower; n'F4 has not been recorded in this form.
Formerly considered a synonym of C. tentans Fabricius, the North American material clearly differs genetically from the Palearctic species and was renamed by Shobanov et al. (1999).
C. dilutus and C. pallidivittatus cannot be separated on the basis of the DNA "barcode" sequence of mtCO1, or CytB (Guryev et al. 2001) but can be separated by the sequence of the globin gene gb2β (Martin et al. 2002).
Molecular sequences:
mtCOI sequences in GenBank, Accession nos. AF110160 - AF110162.
mtcytB sequences in Genbank, Accession nos. AF109700 - AF109709.
gb2β sequences in Genbank, Accession nos. AF110173 - AF110174.
See also C. pallidivittatus and C. tentans [ Return to Index| Go to References ]